Abdellah Barakate
Abdellah joined The James Hutton Institute in November 2015 and he is a member of the Cell and Molecular Sciences [2] department. Prior to this he worked at the University of Dundee for 18 years after his first Postdoc at Imperial College, London. He was born and educated in Morocco and gained his master’s degree and PhD from the University of Joseph Fourier, Grenoble, France.
Abdellah has 3 decades of experience in Plant Molecular Biology and worked mainly in the areas of cell wall and plant DNA repair and recombination. Over years, he has accumulated huge knowledge of gene manipulation tools including CRISPR-based gene editing. Abdellah loves sharing his knowledge through teaching, collaborations and public outreach.
Current research interests
I joined JHI in November 2015 to work on “meiosis and recombination in large genome crop plants”, funded by a 5-year ERC advanced grant awarded to Prof Robbie Waugh [3]. This project extends my previous work in Prof Claire Halpin’s laboratory [4] on “Meiosis in barley: manipulating crossover frequency and distribution” funded by a BBSRC sLoLa grant.
During meiosis, homologous chromosomes undergo complex and dynamic structural changes resulting in their pairing, synapsis, recombination and segregation. The mechanism that regulates the frequency and distribution of meiotic crossovers (CO) is poorly understood. In grasses, meiotic COs are preferentially formed at sub-telomeric regions resulting in stable linkage blocks in centromic region containing ~30% of genes. This skewed pattern of recombination hampers the breeders’ effort to exploit this important gene pool.
Our aim is to understand the mechanism that governs the formation of meiotic COs in barley and use the gained knowledge to improve crop breeding efficiency. In the current SHUFFLE [5] project, we are using several omics platforms ranging from genomics, transcriptomics to proteomics to study the dynamics of RNA and protein changes during meiosis.
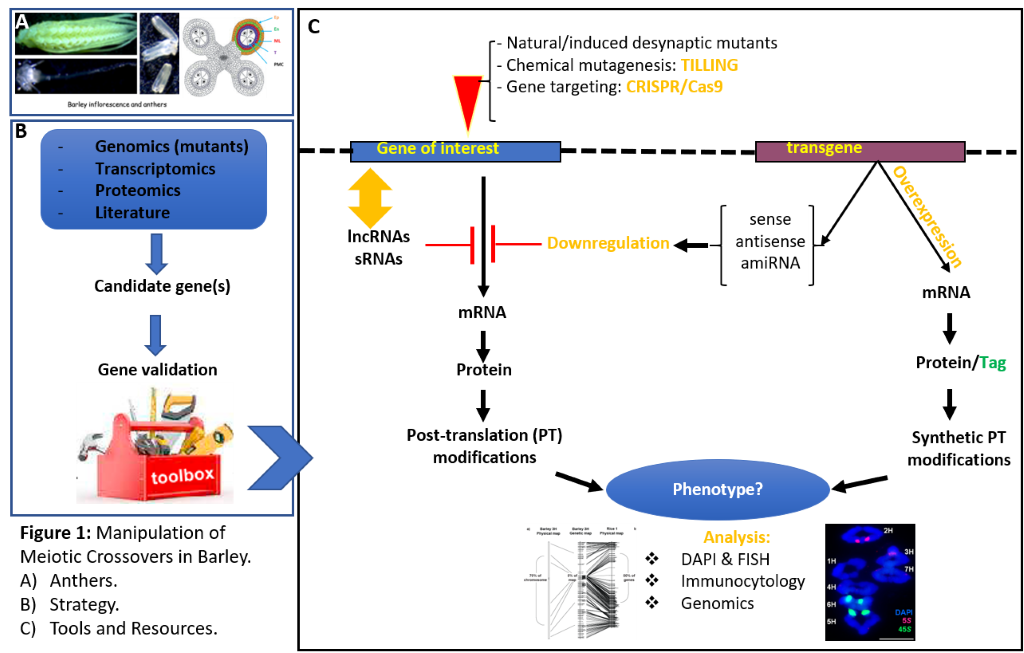 [6]I use molecular biology techniques to study and validate the function of selected genes and Figure 1 summarises my gene manipulation toolbox ranging from RNAi to CRISPR/Cas9 [7].
[6]I use molecular biology techniques to study and validate the function of selected genes and Figure 1 summarises my gene manipulation toolbox ranging from RNAi to CRISPR/Cas9 [7].
I am also interested in studying chromatin epigenetics, non-coding RNAs and post-translational modifications during meiosis. However, these targeted analyses necessitate isolation of large numbers of meiocytes that are embedded in complex structures like anthers. In order to solve this problem, I adapted the INTACT [8] technology (isolation of nuclei tagged in specific cell types) that will allow us to quickly harvest large numbers of meiotic nuclei directly from barley inflorescences.
Collaborations
Currently collaborating with:
- Dr Stefan Heckmann [9], IPK on barley meiotic mutants.
- Dr Jennifer Stephens [10] for improving barley and potato transformations.
- with nine local JHI and University of Dundee groups and ten other international teams on CRISPR/Cas9. I developed user friendly CRISPR/Cas9 vectors for both monocotyledon and dicotyledon plants.
- with Dr Stefania Giacomello [11] and Prof Joakim Lundeberg [12], SciLifeLab, Stockholm to study spatial transcriptome of barley anthers and spikes.
Measures of esteem
- I have reviewed several international and UK research grant proposals.
- I review papers and reviews for several journals [13].
- I have been invited to give talks on lignin biosynthesis, TILLING and CRISPR at local, national and international conferences and workshops.
- I also gave invited lectures to 3rd year students at the University of Dundee on plant transformations and CRISPR.
Employment history
- Nov 2015 – Present: Plant Molecular Geneticist, JHI
- Apr 2015 – Oct 2015: Casual Researcher, University of Dundee.
- Jun 2014 – Mar 2015: Temporary Researcher, JHI.
- Aug 1996 – May 2014: Senior Postdoctoral Researcher, University of Dundee.
- Apr 1993 – Jul 1996: Postdoctoral Researcher, Imperial College, London.
Education and professional qualifications
- October 1989 – December 1992: PhD on pollen development, University Joseph Fourier, Grenoble, France.
- Sep 1988 – Jun 1989: Msc in Molecular and Cell Biology, University of Joseph Fourier, Grenoble, France.
- Summer 1988: Intensive course in Molecular Biology, Rabat, Morocco.
- Sep 1984 – Jun 1988: Bsc in plant biology, biochemistry, microbiology, and genetics.
- Oct 1977 – Jun 1984: Secondary school, Baccalaureate, Beni Mellal, Morocco.
Past research
After my PhD I have worked in two main areas of research:
- Lignin biosynthesis with Prof Claire Halpin [4]
-
Plant DNA repair and recombination:
- My first Postdoc on using tomato golden mosaic virus-based vectors for gene targeting in plants was with Prof Conrad Lichtenstein, Imperial College, London.
-
I then continued this area by applying for funding with Prof Claire Halpin to study intrachromosomal recombination in somatic and meiotic recombination leading to our work on meiosis in barley.
Bibliography
Links:
[1] https://orcid.org/0000-0002-7628-6741
[2] https://www.hutton.ac.uk/research/departments/cell-and-molecular-sciences
[3] https://www.hutton.ac.uk/staff/robbie-waugh
[4] https://www.lifesci.dundee.ac.uk/people/claire-halpin
[5] https://youtu.be/XyMhyeWMZl4
[6] https://www.hutton.ac.uk/sites/default/files/images/research/gene-manipulation-toolbox2.png
[7] https://www.scientificamerican.com/video/crispr-cas9-hd-final/
[8] https://www.youtube.com/watch?v=mazTODTkJzg
[9] https://www.ipk-gatersleben.de/en/independent-research-groups/meiose/
[10] https://www.hutton.ac.uk/staff/jennifer-stephens
[11] https://www.spatialresearch.org/research-giacomello-lab/
[12] https://www.scilifelab.se/researchers/joakim-lundeberg/
[13] https://publons.com/researcher/1181842/abdellah-barakate/metrics/