Science departments
The James Hutton Institute’s commitment to diverse research is channelled through five specialised departments. Each department is a hub of scientific talent, merging expertise and skills to drive innovation and understanding.
From pioneering genetic enhancement in crops, to exploring the intricacies of plant biology, social sciences, computational analysis, and the fascinating world of fungi, plants and soils. Our teams spearhead cutting-edge research across a spectrum of disciplines.
Cell and Molecular Sciences
Cell and Molecular Sciences (CMS) is based at the Dundee site and comprises more than 90 plant scientists. With research specialisms in cell and molecular biology, genomics, genetics, pathology and physiology.
A major research focus is on the genetic improvement of cereals, potatoes and soft fruit crops. With respect to yield and quality, resource use efficiency and pest and disease resistance.
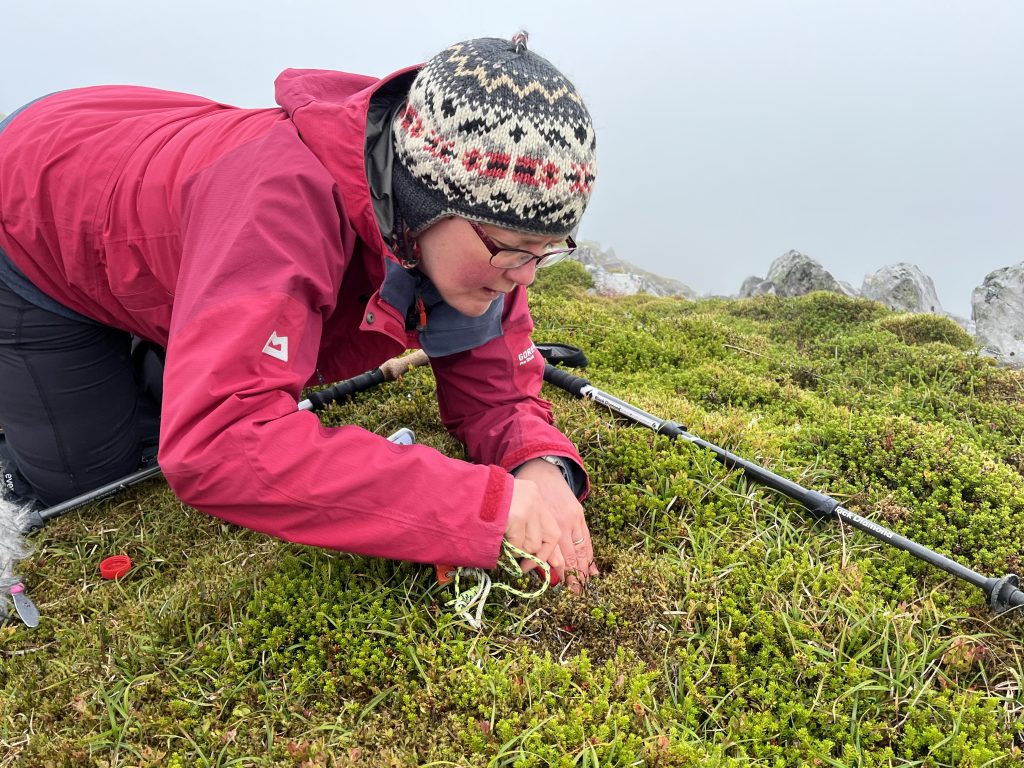

Ecological Sciences
Ecological Sciences (ES) is a multi-disciplinary collective of over 65 researchers. With a unique breadth of scientific expertise, skills and knowledge in the ecology, physiology and systematics of microbes, lichens, fungi, plants, soils, and invertebrates.
Our collective expertise and knowledge is deployed globally across a range of systems. Our dynamic department maintains a large post-graduate student cohort affiliated with universities around the world.
Environmental and Biochemical Sciences
Environmental and Biochemical Sciences (EBS) is a department of over 80 scientists. It spans a range of disciplines encompassing entirely laboratory-based work to ecosystem-scale research across landscapes and rivers.
The breadth and diversity of our work includes the metabolomics of food crops, their chemical makeup and control of this through plant breeding. Our work on soils covers not only their distribution and classification but also function, composition and physical properties; in both natural and managed systems.


Information and Computational Sciences
The Information and Computational Sciences (ICS) department brings together an exceptional combination of scientific skills and expertise.
This ranges from genome scale bioinformatics to the modelling of soil and climate information on a geographical scale. The breadth of skills and resources in ICS provides the James Hutton Institute with a unique capacity to rise to the challenges that come from the new high-throughput data generation technologies that are revolutionising genome, soils, and climate analysis.
Social, Economic and Geographical Sciences
The Social, Economic and Geographical Sciences (SEGS) department represents the fields of economics, geography, politics, sociology, anthropology, psychology and everything in between.
Our mission is to be internationally recognised for the quality, relevance and impact of our social science research connecting land, people and the environment.

